Tutorial¶
The PETGEM tutorial contains a small collection of programs which demonstrate main aspects of the PETGEM work-flow. Each example has the following structure:
Readme.txt. Short description about what the example does.params.yaml. Physical parameters for the 3D CSEM/MT survey.petsc.opts. Parameters for PETSc solvers.Data for modeling (mesh, conductivity model and receivers list)
The data for simple cases modelling are freely available in the Download section.
Basic notions¶
The use of PETGEM can be summarized in three steps: pre-processing, modelling and
post-processing. The pre-processing and modelling phases requires parameter files
where the physical conditions of the 3D CSEM/MT model are defined. In the
params.yaml the 3D CSEM survey is described using several keywords
that allow one to define the main physical parameters and necessary file
locations. In sake of simplicity and in order to avoid a specific parser, the
params.yaml file is defined as Python
dictionaries. Furthermore, the syntax of the params.yaml file is very simple yet powerful. As consequence, the
dictionary names and his key names MUST NOT BE changed. See Preprocessing
parameters file description and Modelling parameters file description
in Manual section for a full explanation of those keywords.
For a general 3D CSEM/MT survey, the PETGEM work-flow can be summarize as follows:
Following the contents of the
params.yamlfile, a set of data are preprocessed (mesh, conductivity model and receivers positions) by thekernel.pyA problem instance is created
The problem sets up its domain, sub-domains, source, solver. This stage include the computation of the main data structures
Parallel assembling of
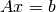 .
.The solution is obtained in parallel by calling a
ksp()PETSc object.Interpolation of electromagnetic responses & post-processing parallel stage
Finally the solution can be stored by calling
postprocess()method. Current version support hdf5.
Based on previous work-flow, any 3D CSEM/MT modelling requires the following input files:
A mesh file (current version supports Gmsh meshes)
A conductivity/resistivity model associated with the materials defined in the mesh file
A list of receivers positions in hdf5 format for the electromagnetic responses post-processing
A
params.yamlfile where are defined the 3D CSEM/MT parametersA
petsc.optsfile where are defined options for PETSc solversA
kernel.pyscript which manage the pre-processing and modelling tasks respectively
Running a simulation¶
This section introduces the basics of running PETGEM on the command line. Following command should be run in the top-level directory of the PETGEM source tree.
PETGEM kernel is invoked as follows:
$ mpirun -n MPI_tasks python3 kernel.py -options_file path/petsc.opts path/params.yaml
where MPI_tasks is the number of MPI parallel tasks, kernel.py is
the script that manages the PETGEM work-flow, petsc.opts is the
PETSc options file and params.yaml
is the modelling parameters file for PETGEM.
A template of this file is included in examples/
of the PETGEM source tree. Additionally, a freely available copy of this file
is located at Download section.
See Model parameters file section for more details about
params.yaml file.
Model parameters file¶
By definition, any 3D CSEM/MT survey should include: physical parameters, a mesh
file, source and receivers files. These data are included in the
params.yanl file. Additionally, options for
PETSc solvers are defined in a
petsc.opts file.
In order to avoid a specific parser, params.yaml file is imported by
PETGEM as a Python dictionary. As consequence, the dictionary name and his key names
MUST NOT BE changed.
A glance of params.yaml file is the following:
###############################################################################
# PETGEM parameters file
###############################################################################
# Model parameters
model:
mode: csem # Modeling mode: csem or mt
csem:
sigma:
#file: 'my_file.h5' # Conductivity model file
horizontal: [1., 0.01, 1., 3.3333] # Horizontal conductivity
vertical: [1., 0.01, 1., 3.3333] # Vertical conductivity
source:
frequency: 2. # Frequency (Hz)
position: [1750., 1750., -975.] # Source position (xyz)
azimuth: 0. # Source rotation in xy plane (in degrees)
dip: 0. # Source rotation in xz plane (in degrees)
current: 1. # Source current (Am)
length: 1. # Source length (m)
mt:
sigma:
file: 'my_file.h5' # Conductivity model file
#horizontal: [1.e-10, 0.01] # Horizontal conductivity
#vertical: [1.e-10, 0.01] # Vertical conductivity
frequency: 4. # Frequency (Hz)
polarization: 'xy' # Polarization mode for mt (xyz)
# Common parameters for all models
mesh: examples/case_CSEM/DIPOLE1D.msh # Mesh file (gmsh format v2)
receivers: examples/case_CSEM/receiver_pos.h5 # Receiver positions file (xyz)
# Execution parameters
run:
nord: 1 # Vector basis order (1,2,3,4,5,6)
cuda: False # Cuda support (True or False)
# Output parameters
output:
vtk: True # Postprocess vtk file (EM fields, conductivity)
directory: examples/case_CSEM/out # Directory for output (results)
directory_scratch: examples/case_CSEM/tmp # Directory for temporal files
A template of this file is included in examples/
of the PETGEM source tree. Additionally, a freely available copy of this file
is located at Download section. Furthermore, in
Running a simulation section of the PETGEM Manual is included
a deep description about this file.
Visualization of results¶
Once a solution of a 3D CSEM/MT survey has been obtained, it should be post-processed by using a visualization program. PETGEM does not do the visualization by itself, but it generates output file (hdf5 format is supported) with the electromagnetic responses (Ex, Ey, Ez, Hx, Hy, Hz) for CSEM and the apparent resistivities and phases for MT. It also gives timing values in order to evaluate the performance.

